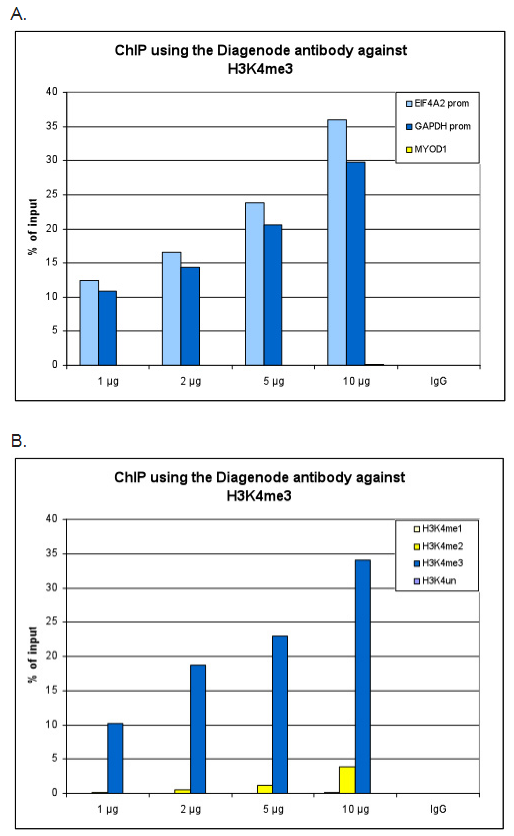
ChIP assays were performed using human HeLa cells, the Bioss antibody against H3K4me3 (Cat. No. bs-53103R) and optimized PCR primer pairs for qPCR. ChIP was performed using sheared chromatin from 1 million cells. The chromatin was spiked with a panel of in vitro assembled nucleosomes, each containing a specific lysine methylation (SNAP-ChIP K-MetStat Panel, Epicypher). A titration consisting of 1, 2, 5 and 10 \u00b5g of antibody per ChIP experiment was analyzed. IgG (2 \u00b5g\/IP) was used as a negative IP control. Figure A. Quantitative PCR was performed with primers specific for the promoter of the active GAPDH and EIF4A2 genes, used as positive controls, and for the inactive MYOD1 gene, used as negative control. The graph shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis). These results are in accordance with the observation that trimethylation of K4 at histone H3 is associated with the promoters of active genes Figure B. Recovery of the nucleosomes carrying the H3K4me1, H3K4me2, H3K4me3 modifications and the unmodified H3K4 as determined by qPCR. The figure clearly shows the antibody is very specific in ChIP for the H3K4me3 modification. At higher concentrations, some H3K4me2 is also precipitated.